现期刊物2026
卷册: 16, 期号: 10
生物化学
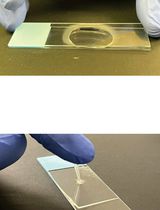
PEPTERGENT: A Peptide-Based Reagent for Detergent-Free Extraction of Membrane Proteins and Purification of Membrane Proteomes
PEPTERGENT:一种用于无去污剂提取膜蛋白及纯化膜蛋白组的多肽试剂
生物信息学与计算生物学
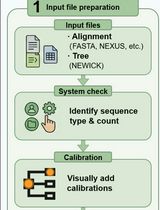
A Step-by-Step GUI-Based Protocol for Molecular Dating Analysis Using PhyloSuite v2
基于 PhyloSuite v2 图形界面的分子定年分析分步操作流程
生物物理学
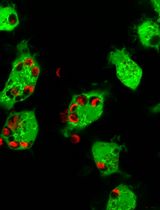
Using Single-Particle Fluorescence Microscopy to Quantify Substrate Binding of Peptidoglycan-Modification Enzymes
利用单颗粒荧光显微成像定量分析肽聚糖修饰酶的底物结合
细胞生物学
Isolation and Biophysical Characterization of Extracellular Vesicles Released by Myocytes
肌细胞来源细胞外囊泡的分离与生物物理特性分析
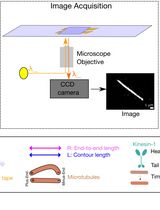
Quantification of Spatial Patterns of Microtubule Transport by Kinesin-1 Head and Tail
驱动蛋白-1头部与尾部介导的微管运输空间模式定量分析
免疫学
A Versatile In Vitro Quantitative Assay for Macrophage Efferocytosis in Diverse Research Applications
适用于多种研究场景的巨噬细胞胞葬作用体外定量检测方法
微生物学
An In Vitro A-431 Epithelial Cell Infection Model for Studying Fungal Pathogenicity and Immune Responses Associated With Vulvovaginal Candidiasis
用于研究外阴阴道念珠菌病相关真菌致病性与免疫反应的 A-431 上皮细胞体外感染模型
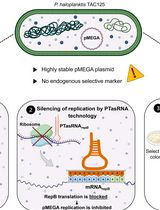
Plasmid Curing of Pseudoalteromonas haloplanktis TAC125 Using Homologous Recombination and PTasRNA Gene Silencing
通过同源重组和PTasRNA基因沉默实现Pseudoalteromonas haloplanktis TAC125质粒清除
神经科学
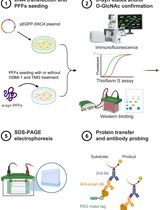
Chemoenzymatic Labeling Method for Detection of O-GlcNAcylated α-Synuclein Proteins by Western Blot
利用化学酶法标记和蛋白质印迹检测 O-GlcNAc 修饰的 α-突触核蛋白
植物科学
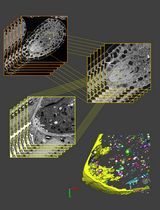
3D Reconstruction of Mature Arabidopsis Ovules Using FIB-SEM to Study Filiform Apparatus Morphology
利用 FIB-SEM 三维重建成熟拟南芥胚珠以研究丝状器形态
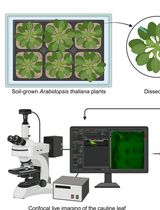
Analysis of Cauline Leaf Development in Arabidopsis thaliana Using Time-Lapse Confocal Microscopy
利用延时共聚焦显微成像分析拟南芥茎生叶发育过程
Biolayer Interferometry (BLI) to Quantify RALF1–Pectin Interactions
利用生物层干涉技术定量分析 RALF1 与果胶的相互作用